Technological progress
Due to rapid advances in the development of state-of-the-art technologies and informatics tools, which were originally used to generate and process biological datasets in biomedicine, the new methods are now being successfully applied in other business sectors as well. Omics technologies provide a holistic view of the molecular characteristics and processes of a cell, tissue or complete organism. “Omics” therefore refers to novel overall concepts for analyzing the complete genetic or molecular profile of organisms using integrative pipelines. These reveal how complex interactions between genes and molecules influence phenotype, such as a patient’s disease symptoms. Extensive technological improvements regarding next-generation sequencing, microarray, mass spectrometry, and nuclear magnetic resonance have fundamentally changed biological and biomedical research in recent years. Omics data analysis is an expanding field, offering new perspectives not only to the medical field through the creation of ever new statistical methods.
Examples on different omics data types:
- Genomics: totality of all genes of an organism
- Proteomics: totality of all proteins produced in an organism.
- Transcriptomics: totality of all RNA molecules
- Pharmacogenomics: The Influence of Genetic Modifications on the Effect of Drugs
Compared to single omics analyses, multi-omics queries provide researchers with more comprehensive information, from the original cause of disease (genetic, environmental, or developmental) to functional effects or relevant interactions.
Computational Biology
Bioinformatics uses computer-based tools to manage and analyze biological data from large data sets. The integrative approach of combining such tools with corresponding databases is called computational biology. It allows the creation of entire maps of cellular and physiological signaling pathways. Bioinformatics methods enable the holistic representation of a cell or an organism as well as predictions of highly complex systems such as phenotypes or networks of cellular interactions (Fig. 1).
The convergence between omics and bioinformatics represents the basis of systems biology. This area of study generates model organisms to gain a deeper understanding of complex biological interactions within cells or tissues at the gene, protein, or metabolite level. [2,5] But as the generation of such datasets takes less and less time and cost, the integration of omics data offers both groundbreaking opportunities and immense challenges for biologists, biostatisticians, and biomathematicians.
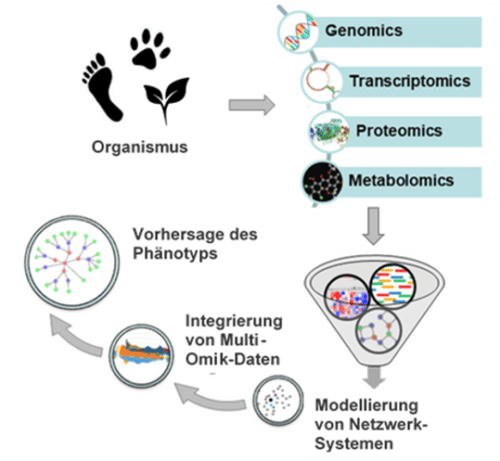
Figure 1; Holistic understanding through integrated omnik analyses
Challenge
Today, there are countless web-based solutions for storing, analyzing and sharing data. Examples of commonly known portals with download capability of omics data are Gene Expression Omnibus, ArrayExpress, Expression Atlas or Ensembl. Other portals additionally provide options for analyzing the corresponding data in interactive and multi-layered levels, such as NCBI, UCSC, or the Human Brain Atlas. Due to the ever-increasing number of bioinformatics tools, special platforms for listing the tools have also been created to ensure their effective use. EBI, for example, provides an extensive portal with a large number of databases and tools sorted by topic. OMICtools, on the other hand, offers a library of specialized software solutions, databases and platforms for processing and analyzing large amounts of data.
But despite the existence of countless omics tools for whole network analysis, pathway representation, genome alignments, visualization, and many other uses, hardly any of these solutions can integrate 3 or more different omics datasets. Thus, managing and integrating multidimensional data remains difficult, in part because each omics analysis can generate tera- or peta-byte-sized files on a daily basis. Currently, the biggest challenges are the biological and technical differences in terms of data preparation, storage and processing, scientific contextualization, statistical validation, and computational power. There is also a lack of stable pipelines that can integrate additional data types. Innovative approaches to incorporate omics data are urgently needed, especially in the fields of nutritional sciences, microbiome analyses, genotype-phenotype interaction, systems biology, and disease research.
Importance of Omik Tools in Medicine
Extensive pofiling now provides deep insights into the development of serious and chronic diseases such as cancer, cardiac or infectious diseases, facilitates the search for diagnostic markers or efficient novel therapies and allows the prediction of therapeutic success. The identification of predictive and early diagnostic markers is of great advantage especially in cancer therapy, as some cancers are diagnosed only at an advanced stage and the chances of survival are consequently low.
Integrative multi-omics analyses are designed to provide a holistic view of disease mechanisms that interfere with the execution of normal cellular functions and lead to disease progression and drug resistance. However, while computer-assisted analyses of single omics datasets are now well established, approaches to integrating multi-omics data are still far from standard. To keep up with the enormous speed of data generation and growth of biological knowledge, existing methods need to be extended or generalized. New omics tools must also be able to handle the complexity and multi-layered levels of information available.
Importance of Omik Tools in other sectors
Not only the medical field benefits from specialized bioinformatics tools, but also other important economic sectors such as agriculture or plant and animal science. Honey bees, for example, are irreplaceable pollinators for global agriculture. The health of honey bees has deteriorated rapidly in recent decades, with beekeepers losing more than a quarter of their colonies per year in some cases since 2007. The reasons for bee mortality are complex, independent of time and place, and very difficult to define.
The Canadian BeeCSI project, launched in 2018, now aims to use genomic tools to develop a new platform for assessing and diagnosing current bee health based on stress factors and corresponding biomarkers. Two specialists from York University and the University of British Columbia are leading a national team of researchers, bioinformaticians, beekeepers and diagnostic laboratories. BeeCSI is expected to bring about industrial modernization in Canada in the form of this new tool, which allows rapid assessment of the health status of bee colonies. This enables beekeepers to take appropriate measures against bee mortality at an early stage. The $10 million project is funded by Genome Canada and Ontario Genomics.
Perspektives
Studies in genomics, transcriptomics, and proteomics have significantly influenced our understanding of cellular complexity and heterogeneity. Due to steadily falling costs of omics analyses, even more types of data can now be integrated in the medical field to create individualized treatment plans. However, the road to standardization is a long one: generating omics data across scales, developing new analytical methods, adapting these methods to specific diseases, and replicating these processes for additional diseases are fundamental tasks in omics research. However, these cannot be handled by one team alone. To ensure the active flow of information between different disciplines, coordinated action must be taken by many teams with appropriate expertise. This may eventually enable standardization of data formats and pipelines for integrated multi-omics analysis.



